标准培养限制有几种生理和病理过程,其中细胞经历体积变化。然而,没有可靠的方法可用于实时准确测量贴壁细胞的体积。 细胞体积测定的优势细胞在光学透明的腔室中培养,该腔室能够间准确确定它们的体积,并并行跟踪导致体积变化的生化过程,例如离子泵的激活。 例子从间期到 Raji 细胞有丝分裂的体积跟踪 [2]。
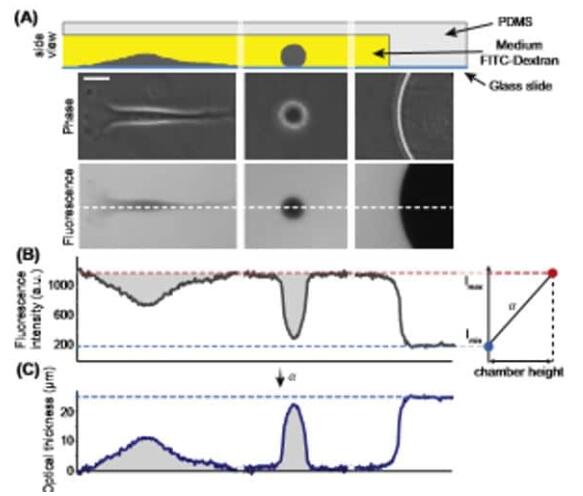
(A) 将细胞置于由柱设定的校准高度的聚(二甲基硅氧烷)室中,在补充有FITC葡聚糖的培养基中。下图:细胞在荧光图像上排除荧光(比例尺20 mm)。 (B) 对应于(A)中虚线的荧光文件:荧光强度的大值和小值分别对应于腔室大高度(背景)和零高度(柱)。右图:这些值用于校准信号并计算细胞的光学厚度。 (C) 后,通过对细胞面积上的总荧光强度进行积分来获得细胞体积。 (A) HeLa 细胞的体积轨迹。开始和结束时的两个体积超调对应于有丝分裂中的瞬时体积增加,一个对应于母细胞,二个对应于子细胞。 (B) (A) 中使用 FXm 的细胞的原始荧光图像。比例尺 50 毫米。
参考[1] Zlotek-Zlotkiewicz, E. 等人。(2015)。细胞生物学杂志,211 (4),765–774。 [2] Cadart, C 等人。(2017)。细胞生物学方法,139,103-120。
提供实现工具细胞限制器-cell confiner(Cell confinement) 我们的cell Confiner是一种多功能设备,可通过对细胞应用定义明确的约束条件来研究细胞力学。限制方法基于将细胞固定在两个平行表面之间,从而实现均匀且定义明确的物理参数,例如细胞几何形状和环境弹性。此外,可以使用高分辨率显微镜对受限的细胞进行成像,因为该设备是光学透明的,并将细胞保持在焦平面上。 细胞被均匀地限制/压缩在两个亚微米分辨率的两个平行表面之间。两个表面之间的空间由微型PDMS支柱控制。 微型支柱在载玻片上制造,载玻片连接到PDMS活塞(吸盘)上。 活塞由真空泵控制,因此限制区的高度也受到控制。不同的限制高度(例如1um – 300um),允许长期细胞培养和细胞增殖,同时保持对封闭的完美控制
与高分辨率光学显微镜系统兼容,可以处理足够多的细胞以进行完整的基因表达分析,可与生物功能化的微结构化底物和/或不同的基质(几何形状控制)结合使用
可以与凝胶结合(硬度控制),兼容任何细胞培养底物(培养皿至96孔板)。 产品特性: >定义细胞的厚度和形状 用正确的限制滑片控制细胞的厚度 >同时进行多个实验 能够研究不同的细胞或同时应用不同的限制条件 >适用于高分辨率显微镜 光学透明的材料和紧凑的设计可实现高分辨率显微镜 >控制限制速度 通过真空泵j确控制限制速度 >可逆限制:限制后取回您的细胞 由于细胞的非破坏性方法,可以进行分子分析 >与您自己的实验兼容 该限制器是一种小型设备,直接放置在您的细胞培养液顶部 技术方案图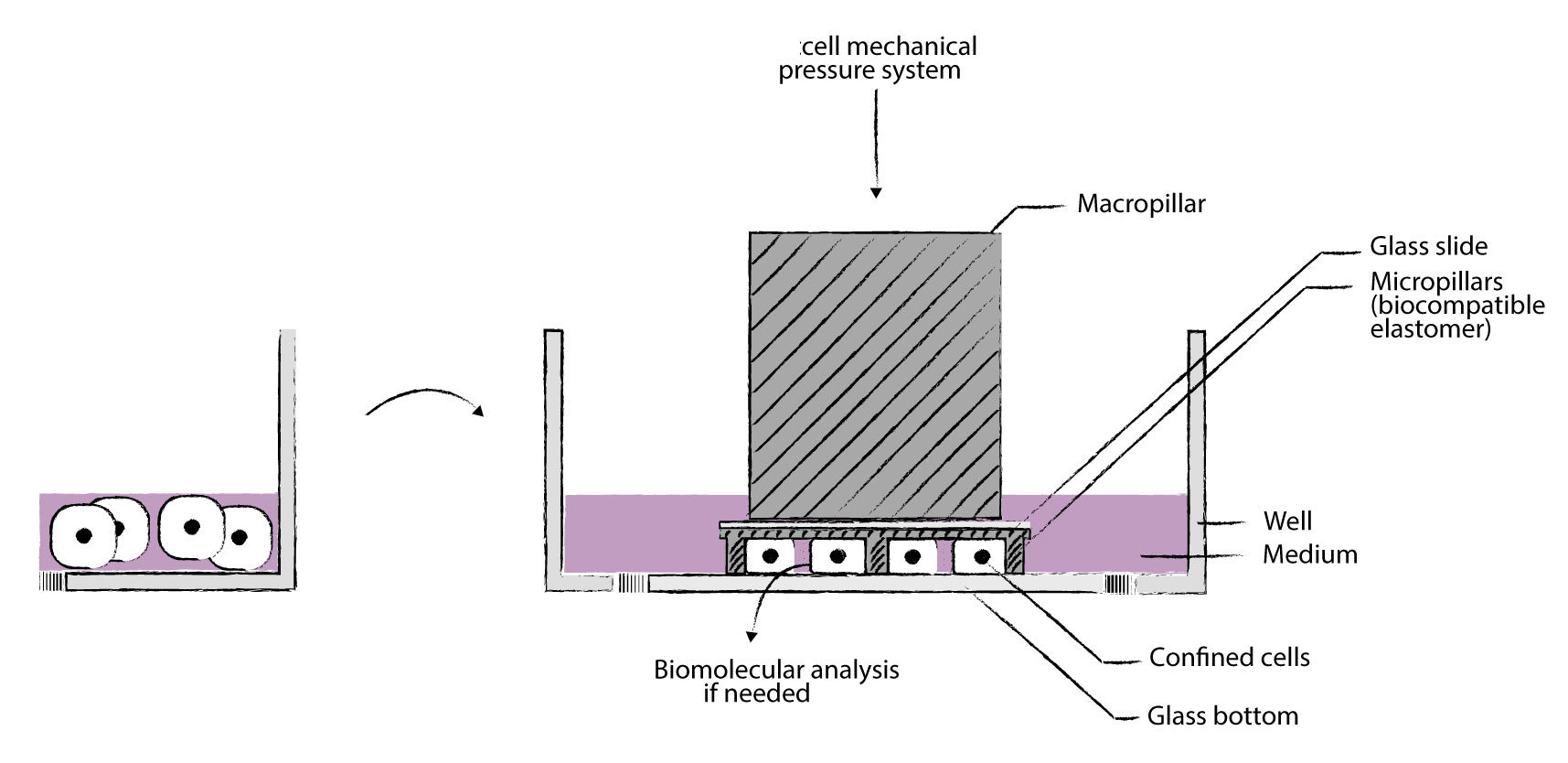
三种产品形式典型应用:>癌症浸润测定:迁移行为和迁移转变的量化 >癌症侵袭性测定:体细胞或癌细胞的收缩力定量 >内吞作用测定:更好地观察膜发生的事件 >胞吐法测定:更好地观察在顶端膜发生的事件 >吞噬功能失调:机制的表征 >孔中的免疫系统:非粘附免疫细胞的二维迁移和相互作用 >免疫细胞相互作用:非贴壁免疫细胞的2D相互作用 >有丝分裂组装测定:有丝分裂纺锤体疾病的定量 >定量细胞迁移测定:细胞迁移特性的快速,精细分析 >癌症研究 转移细胞的迁移 转移中的细胞收缩 DNA DSB修复(机械诱导) 基因组不稳定(细胞分裂) 分离共培养 >免疫学 免疫细胞迁移 非粘附细胞的成像 >器官生理学 癌细胞迁移 具有硬度控制的细胞区分 伤口愈合 分离共培养 细胞压缩反应 >罕见疾病 细胞核完整性 >老化 细胞核完整性 自噬相关疾病 >观测化 非粘附细胞的成像 细胞器的平面成像 >基础研究 细胞体积(细胞周期) 细胞机械力刺激反应 二维心肌细胞成熟测定
二维肝小管化验
3D心肌细胞成熟测定
3D肝小管测定
附着球体测定
细胞收缩力测定
细胞迁移测定
细胞核挤压测定
细胞j化
细胞体积测量
趋化性测定
共培养测定
胞吞试验
胞吐法
外泌体测定
片状脂蛋白和丝状体含量测定
活细胞成像
巨噬细胞j化测定
MT依赖性运输测定
神经肌肉连接测定
井中的神经元网络
细胞器定位分析
初次纤毛测定
骨骼肌细胞测定
平滑肌细胞
伤口愈合测定
PUBLICATIONS - Confinement and Low Adhesion Induce Fast Amoeboid Migration of Slow Mesenchymal Cells
Y.-J. Liu, M. Piel, Cell, et al., 2015 160(4), 659-672 - Actin flows induce a universal coupling between cell speed and cell persistence
P. Maiuri, R. Voituriez, et al., Cell, 2015 161(2), 374–386 - Geometric friction directs cell migration
M. Le Berre, M. Piel, et al., Physical Review Letter 2013 111, 198101 - Mitotic rounding alters cell geometry to ensure efficient spindle assembly
O. M. Lancaster, B. Baum, et al., Developmental Cell, 2013 25(3), 270-283 - Fine Control of Nuclear Confinement Identifies a Threshold Deformation leading to Lamina Rupture and Induction of Specific Genes
M. Le Berre, J. Aubertin, M. Piel, Integrative Biology, 2012 4 (11), 1406-1414 - Exploring the Function of Cell Shape and Size during Mitosis
C. Cadart, H. K. Matthews, et al., Developmental Cell, 2014 29(2), 159-169 - Methods for Two-Dimensional Cell Confinement
M. Le Berre, M. Piel, et al., 2014, Micropatterning in Cell Biology Part C, Methods in cell biology, 121, 213-29
References [1] D. Huh, G. A. Hamilton, and D. E. Ingber, “From 3D cell culture to organs-on-chips,” Trends Cell Biol., vol. 21, no. 12, pp. 745–754, 2011. [2] M. Ravi, V. Paramesh, S. R. Kaviya, E. Anuradha, and F. D. Paul Solomon, “3D cell culture systems: Advantages and applications,” J. Cell. Physiol., vol. 230, no. 1, pp. 16–26, 2015. [3] J. W. Haycock, 3D cell culture: a review of current approaches and techniques., vol. 695. 2011. [4] F. Pampaloni, E. G. Reynaud, and E. H. K. Stelzer, “The third dimension bridges the gap between cell culture and live tissue.,” Nat. Rev. Mol. Cell Biol., vol. 8, no. 10, pp. 839–845, 2007. [5] J. Lee, M. J. Cuddihy, and N. A. Kotov, “Three-dimensional cell culture matrices: state of the art.,” Tissue Eng Part B Rev, vol. 14, no. 1, pp. 61–86, 2008. [6] M. Vinci et al., “Advances in establishment and analysis of three-dimensional tumor spheroid-based functional assays for target validation and drug evaluation,” BMC Biol., vol. 10, no. 1, p. 29, 2012. [7] B. A. Justice, N. A. Badr, and R. A. Felder, “3D cell culture opens new dimensions in cell-based assays,” Drug Discov. Today, vol. 14, no. 1–2, pp. 102–107, 2009. [8] I. Meyvantsson and D. J. Beebe, “Cell culture models in microfluidic systems.,” Annu. Rev. Anal. Chem., vol. 1, pp. 423–449, 2008. [9] E. W. K. Young and D. J. Beebe, “Fundamentals of microfluidic cell culture in controlled microenvironments,” Chem Soc Rev, vol. 39, no. 3, pp. 1036–1048, 2010. [10] D. J. Beebe, G. a Mensing, and G. M. Walker, “Physics and applications of microfluidics in biology.,” Annu. Rev. Biomed. Eng., vol. 4, pp. 261–286, 2002. [11] J. El-Ali, P. K. Sorger, and K. F. Jensen, “Cells on chips.,” Nature, vol. 442, no. 7101, pp. 403–411, 2006. [12] J. Guck et al., “Optical deformability as an inherent cell marker for testing malignant transformation and metastatic competence,” Biophys J, vol. 88, no. 5, pp. 3689–3698, 2005. [13] S. K?ster et al., “Drop-based microfluidic devices for encapsulation of single cells.,” Lab Chip, vol. 8, no. 7, pp. 1110–1115, 2008. [14] H. Andersson and A. Van den Berg, “Microfluidic devices for cellomics: A review,” Sensors Actuators, B Chem., vol. 92, no. 3, pp. 315–325, 2003. [15] M. W. Tibbitt and K. S. Anseth, “Hydrogels as extracellular matrix mimics for 3D cell culture,” Biotechnol. Bioeng., vol. 103, no. 4, pp. 655–663, 2009. [16] J. P. Vacanti and R. Langer, “Tissue engineering: the design and fabrication of living replacement devices for surgical reconstruction and transplantation.,” Lancet, vol. 354, p. SI32-I34, 1999. [17] G. S. D. Hetal Patel, Minal Bonde, “Biodegradable polymer scaffolds for tissue engineering,” Trends Biomater. Artif. Organs, vol. 25, no. 1, pp. 20–29, 2011. [18] L. G. Griffith and M. A. Swartz, “Capturing complex 3D tissue physiology in vitro.,” Nat. Rev. Mol. cell Biol., vol. 7, no. 3, pp. 211–24, 2006. [19] D. J. Tobin, “Scaffolds for Tissue Engineering and 3D Cell Culture,” Methods Mol. Biol., vol. 695, no. 2, pp. 213–227, 2011. [20] J. Naranda et al., “Polyester type polyHIPE scaffolds with an interconnected porous structure for cartilage regeneration,” Sci. Rep., vol. 6, no. February, p. 28695, 2016. [21] B. Dhandayuthapani, Y. Yoshida, T. Maekawa, and D. S. Kumar, “Polymeric scaffolds in tissue engineering application: A review,” Int. J. Polym. Sci., vol. 2011, no. ii, 2011. [22] F. J. O’Brien, “Biomaterials & scaffolds for tissue engineering,” Mater. Today, vol. 14, no. 3, pp. 88–95, 2011. [23] A. L. Paguirigan and D. J. Beebe, “Microfluidics meet cell biology: Bridging the gap by validation and application of microscale techniques for cell biological assays,” BioEssays, vol. 30, no. 9, pp. 811–821, Sep. 2008. [24] F.-Q. Nie, M. Yamada, J. Kobayashi, M. Yamato, A. Kikuchi, and T. Okano, “On-chip cell migration assay using microfluidic channels.,” Biomaterials, vol. 28, no. 27, pp. 4017–4022, 2007. [25] A. Valster, N. L. Tran, M. Nakada, M. E. Berens, A. Y. Chan, and M. Symons, “Cell migration and invasion assays,” Methods, vol. 37, no. 2, pp. 208–215, 2005. [26] C. R. Justus, N. Leffler, M. Ruiz-Echevarria, and L. V Yang, “In vitro cell migration and invasion assays.,” J. Vis. Exp., vol. 752, no. 88, p. e51046, 2014. [27] N. Kramer et al., “In vitro cell migration and invasion assays.,” Mutat Res, vol. 752, no. 1, pp. 10–24, 2013. [28] J. W. Hong, V. Studer, G. Hang, W. F. Anderson, and S. R. Quake, “A nanoliter-scale nucleic acid processor with parallel architecture.,” Nat. Biotechnol., vol. 22, no. 4, pp. 435–439, 2004. [29] J. Q. Boedicker, L. Li, T. R. Kline, and R. F. Ismagilov, “Detecting bacteria and determining their susceptibility to antibiotics by stochastic confinement in nanoliter droplets using plug-based microfluidics.,” Lab Chip, vol. 8, no. 8, pp. 1265–1272, 2008. [30] G. Velve-Casquillas, M. Le Berre, M. Piel, and P. T. Tran, “Microfluidic tools for cell biological research,” Nano Today, vol. 5, no. 1. pp. 28–47, 2010. [31] C. R. Terenna et al., “Physical Mechanisms Redirecting Cell Polarity and Cell Shape in Fission Yeast,” Curr. Biol., vol. 18, no. 22, pp. 1748–1753, Nov. 2008. [32] G. Faure-andré, “Regulation of Dendrc Cell Migration by CD74, the MHC Class II–Associated Invariant Chain,” Science (80-. )., vol. 1705, no. December, 2008. [33] S. M. McFaul, B. K. Lin, and H. Ma, “Cell separation based on size and deformability using microfluidic funnel ratchets,” Lab Chip, vol. 12, no. 13, pp. 2369–2376, 2012. [34] S. C. Hur, N. K. Henderson-MacLennan, E. R. B. McCabe, and D. Di Carlo, “Deformability-based cell classification and enrichment using inertial microfluidics.,” Lab Chip, vol. 11, no. 5, pp. 912–920, 2011. [35] H. W. Hou, Q. S. Li, G. Y. H. Lee, A. P. Kumar, C. N. Ong, and C. T. Lim, “Deformability study of breast cancer cells using microfluidics,” Biomed. Microdevices, vol. 11, no. 3, pp. 557–564, 2009.
相关产品: 1、细胞力产生和力传递综合测量分析系统 2、各种生物力学设备
|
